Dockamon – PyRx
Complete molecular docking, virtual screening and ligands scoring software package.
Dockamon – PyRx is a structure-based drug design software package containing two integrated software programs.
PyRx: Molecular docking and virtual screening software.
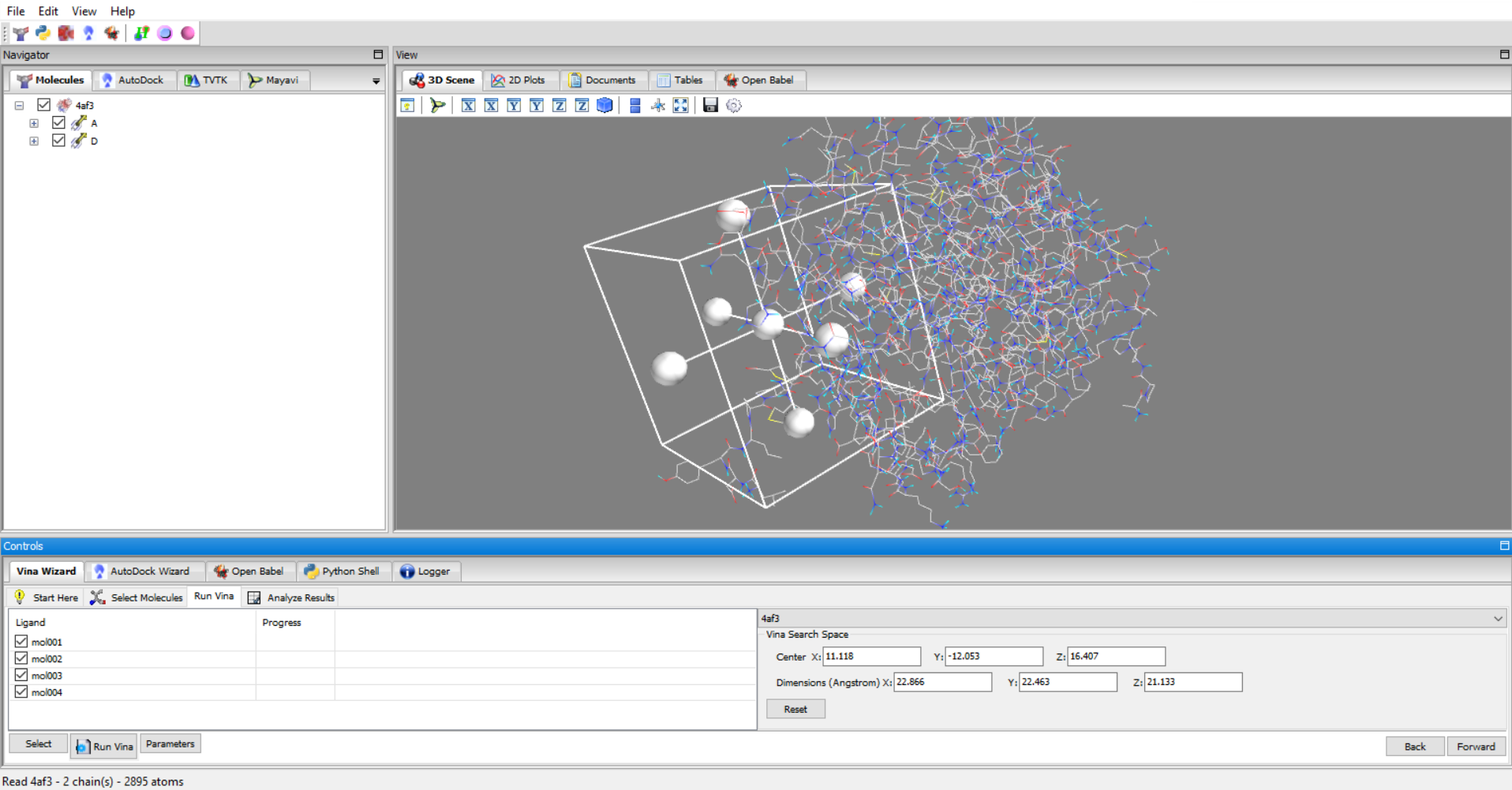
- Molecular Docking: Perform molecular docking using AutoDock and AutoDock Vina software,
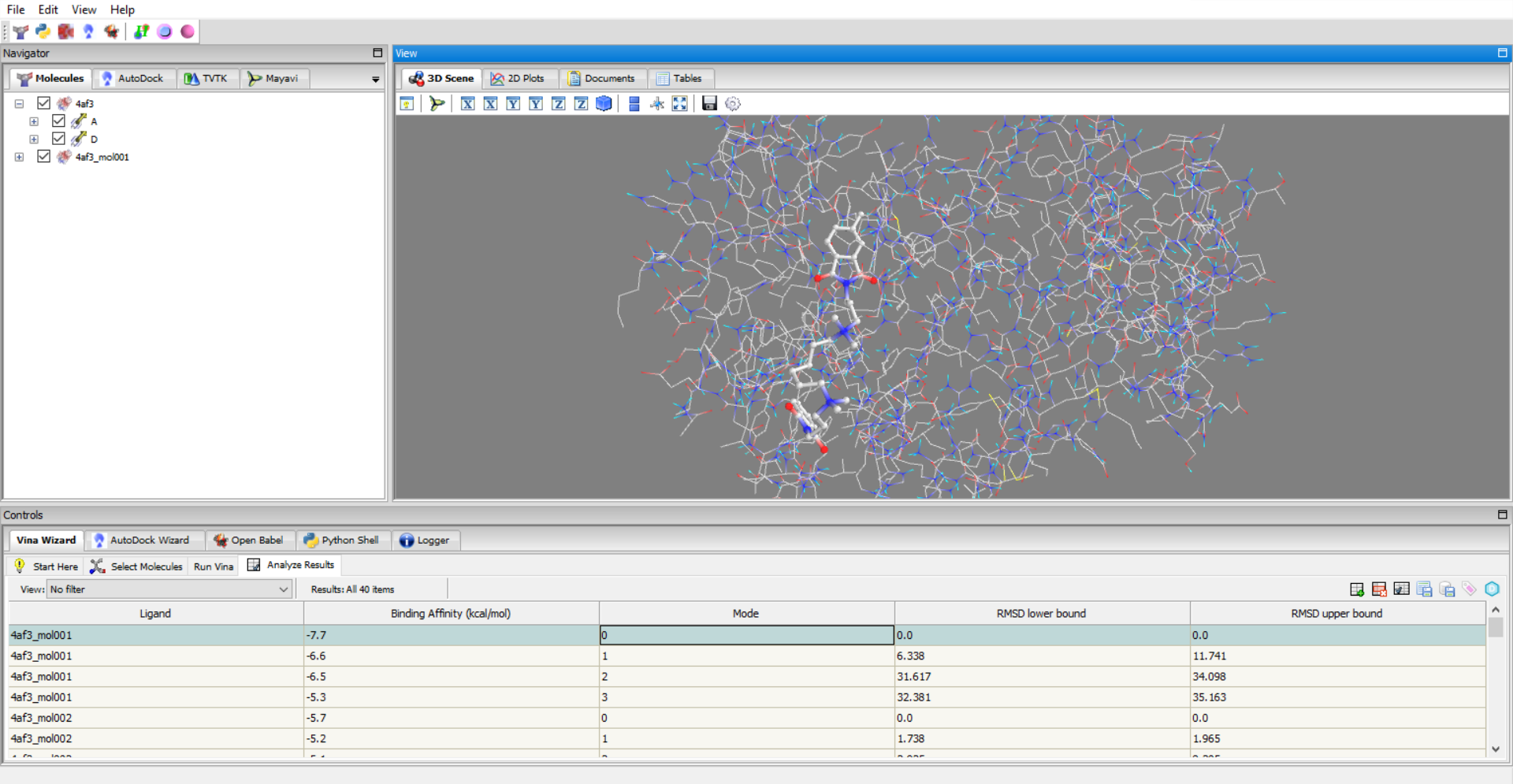
- Virtual Screening: Perform docking on datasets automatically, analyze and export results conveniently.
- Ligands Preparation: Perform energy minimization on ligands and convert to PDBQT format.
- Other Features: Other useful functionalities, including 3D-molecular viewer and the identification of similar compounds in the PubChem database.
Dockamon: Ligands scoring software utilizing machine learning-based scoring functions.
- Ligands Scoring: Score docked poses using a machine learning scoring function including RF-Score V2 and SVM-Score.
- Consensus Scoring: Combine scores from more than one scoring function to achieve better predictions.
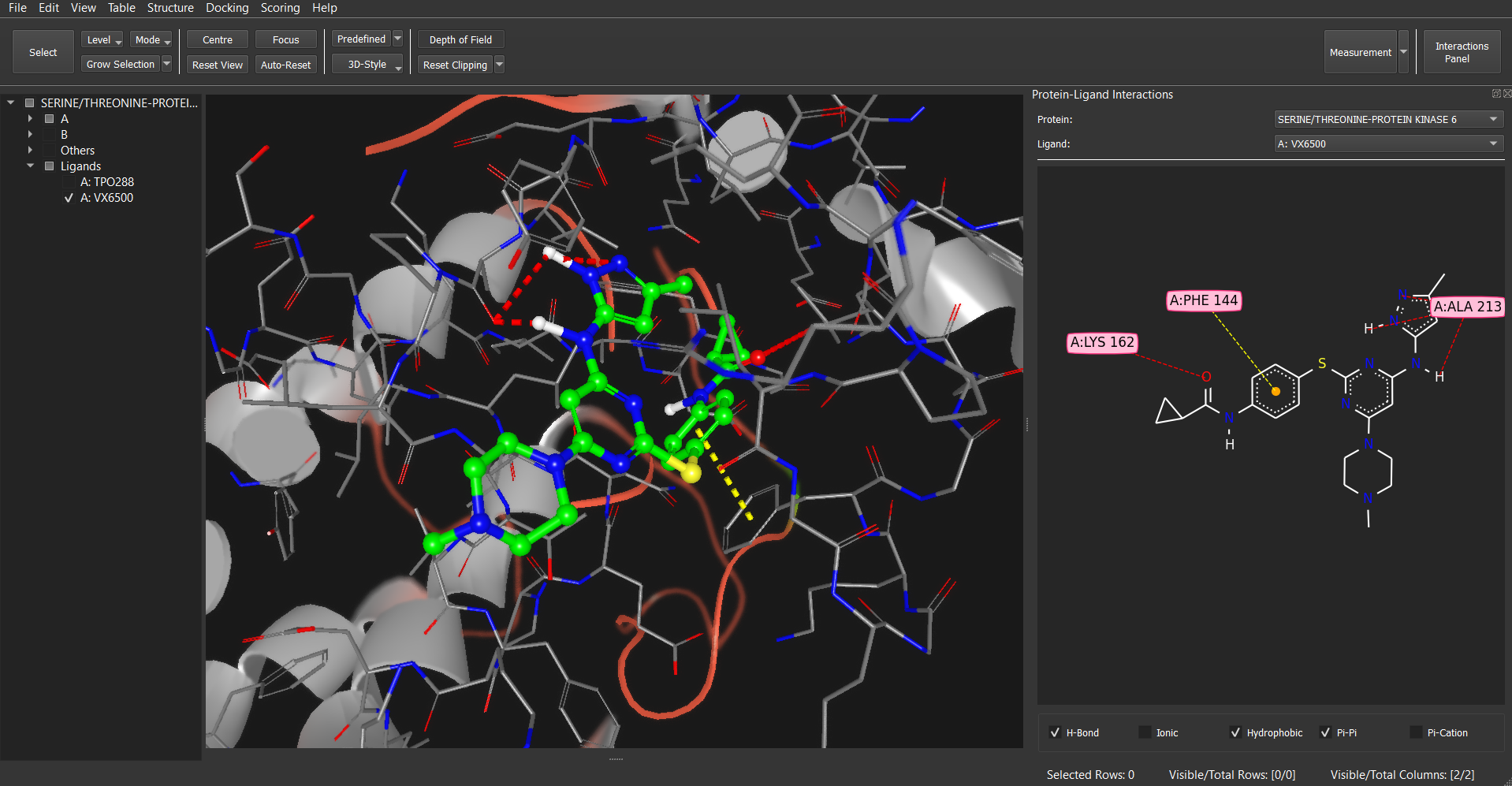
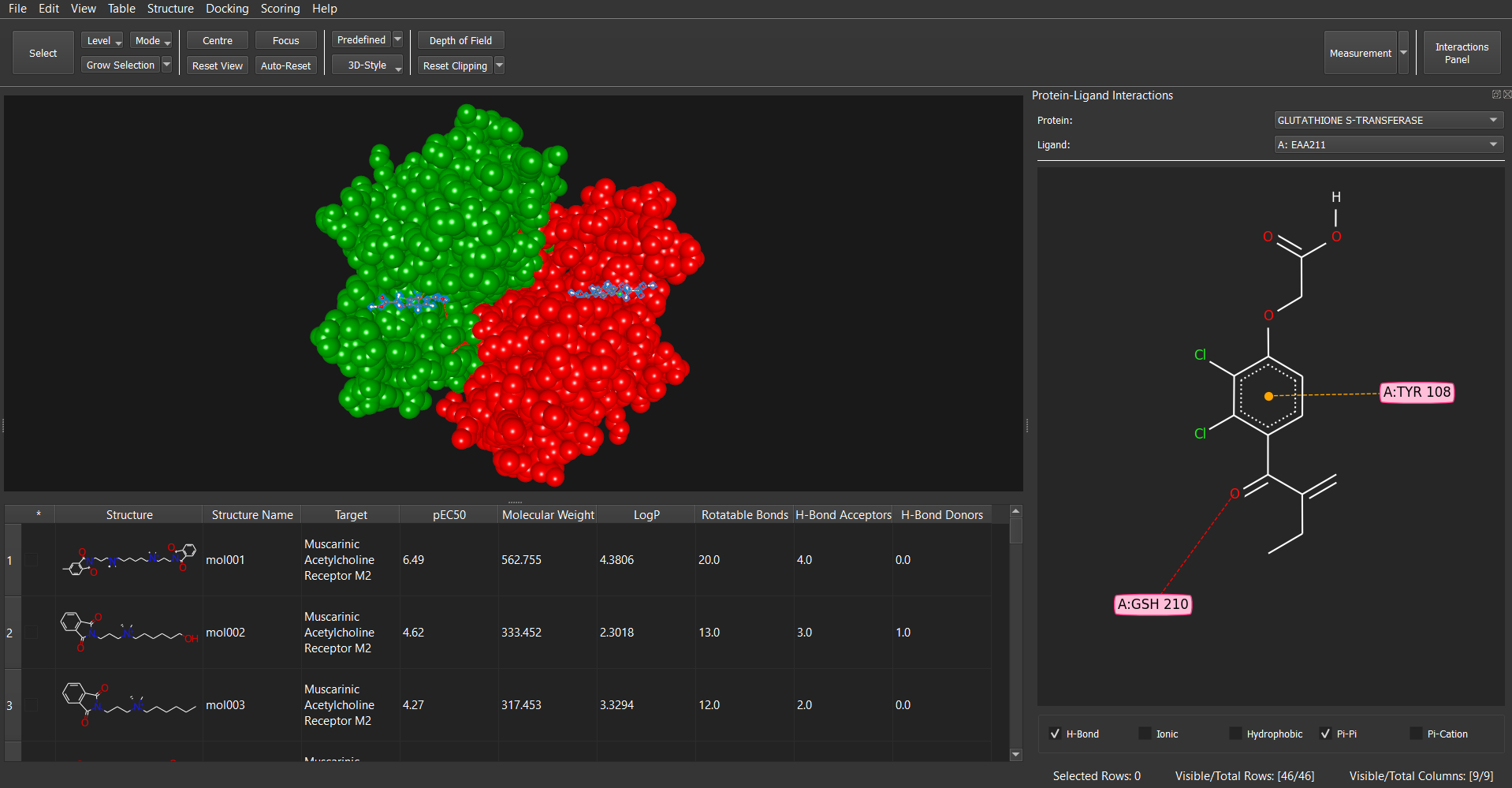
- Protein-Ligand Interactions: Publication quality 2D and 3D interactive visualization of protein-ligand interactions. Customizable interactions options.
- Protein and Ligand Preparation: Prepare protein structures and ligands for docking, scoring or other molecular modeling tasks.
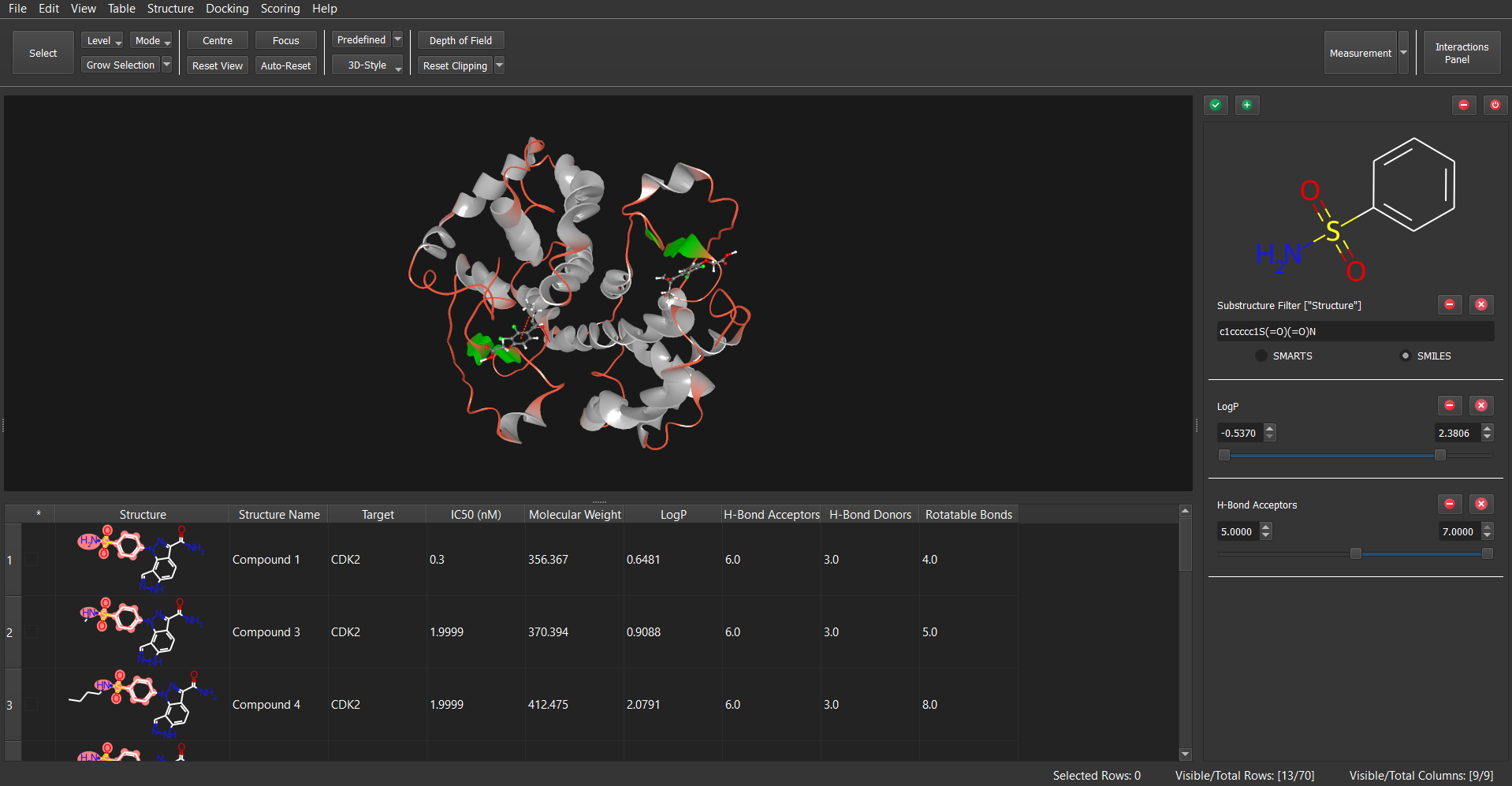
- Filtering & Substructure Matching: SMARTS and SMILES pattern substructure search, numerical and text-based filtering of the data.
- Other Features: Additional functionalities, including chemical spreadsheet capabilities, protein structure management, descriptor calculations, and conformer generation.
Download
Dockamon – PyRx 1.0
$1,394.58 (Coupon: ACADEMIC)
Dockamon – PyRx 1.0
$2,789.16
Use coupon ACADEMIC to get 50% when purchasing. Purchasing the software is facilitated via FastSpring, a trusted platform for software distribution.
If you have any queries or issues, please contact us at: crescent_silico@outlook.com.
PyRx and Dockamon are complementary software that provide powerful structure-based drug design tools. The PyRx software can be used to prepare structures and perform molecular docking using AutoDock and AutoDock Vina. The results of the docking experiment can be directly opened in the Dockamon software.
The Dockamon software is used for scoring ligand docked poses using machine learning scoring functions. The machine learning scoring functions have been demonstrated in the literature to have superior ability to estimate the activity compared to classical scoring functions. For example, the RF-Score V2 scoring function that is implemented in Dockamon, has a Rp value of 0.8 on the Core Set (trained on the Refined Set (v2020)), which is significantly superior compared to the Vina scoring function [1].
Dockamon also offers high-quality 2D and 3D visualization of protein-ligand interactions for publication purposes. These visualizations are fully customizable in terms of visualization settings, and users can customize interaction settings, such as feature definitions and distance cutoffs.
Other functionalities include protein and ligand preparation tasks such as adding hydrogens, assigning ligand bond orders, generating 3D-coordinates, etc. Also, Dockamon provides filtering panel that allows for filtering of structures via substructure, numerical and textual values. Thus, Dockamon can be seamlessly integrated with docking results, enhancing research outcomes by employing machine learning scoring functions and visualizing protein-ligand interactions.
[1] Ballester PJ, Schreyer A, Blundell TL. Does a more precise chemical description of protein–ligand complexes lead to more accurate prediction of binding affinity?. Journal of chemical information and modeling. 2014 Mar 24;54(3):944-55.